Repositories
A Data Repository is a system that stores and publishes Data Files for download. Such repositories have their own architecture, data access controls, data portals and download clients. Generally speaking, there are two types of repositories:
- Cloud - offers facilities for compute and storage
- Non-Cloud - provides storage functionality only
On this page you can find an overview of all of these repositories, their purpose and function, as well as providing links to each repositories' important pages and resources.
All of the ICGC data be searched for using the ICGC Data Portal. Generally the data is divided into projects. Every repository has a repository code / id that is used to identify it in the ICGC API.
Collaboratory
Academic research cloud infrastructure built to house ICGC data.
| Property | Value |
|---|---|
| Name | Cancer Genome Collaboratory |
| Contact | dcc-support@icgc.org |
| Repository type | Cloud |
| ICGC Portal Page | Portal |
| Download Client | Tarball, Docker |
| Repo Code | collaboratory |
Obtaining Data Access
Follow the procedure outlined at the DACO page.
Download Client Operation
To operate the Score download client, follow the instructions here.
AWS
Amazon cloud service containing ICGC data.
| Property | Value |
|---|---|
| Name | ICGC Storage Server (hosted at AWS) |
| Contact | dcc-support@icgc.org |
| Repository type | Cloud |
| ICGC Portal Page | Portal |
| Download Client | Tarball, Docker |
| Repo Code | aws-virginia |
Obtaining Data Access
Follow the procedure outlined at the DACO page.
Download Client Operation
To operate the Score download client, follow the instructions here.
EGA
The European Genome-Phenome Archive (EGA) is co-managed by EBI and CRG. Data can only be downloaded through their EGA download client, but metadata may be viewed on their website. Files are grouped into datasets based on the study they were collected in, and access is granted on a dataset by dataset basis. This repository carries both ICGC and non-ICGC data.
| Property | Value |
|---|---|
| Name | European Genome Archive |
| Contact | helpdesk@ega-archive.org |
| Repository type | Non-Cloud |
| Offical Website | https://ega-archive.org |
| ICGC Portal Page | Portal |
| Download Client | Zipfile |
| Repo Code | ega |
Obtaining Data Access
Follow the procedure outlined at the DACO page. Once approved by ICGC DACO, you will need to contact EGA to have your EGA account set up.
Download Client Operation
To operate the EGA download client, follow the instructions here.
GDC
The Genomic Data Commons is a US government (NIH / NCI) run data repository for cancer genomic information. Notably, the it carries data from The Cancer Genome Atlas (TCGA) and the Therapeutically Applicable Research to Generate Effective Treatments (TARGET). Currently the GDC is the largest single repository of ICGC data. It focuses on studies in the United States.
| Property | Value |
|---|---|
| Name | Genomic Data Commons |
| Contact | support@nci-gdc.datacommons.io |
| Repository type | Non-Cloud |
| Official Data Portal | https://portal.gdc.cancer.gov/ |
| ICGC Portal Page | Portal |
| Download Client | Download the client here |
| Repo Code | gdc |
Obtaining Data Access
To obtain access you must have an eRA commons account, and you must have dbGap access to the data on the GDC you are interested in. Talk to your team leader if you do not have this access. Once this account is set up, you can log in to the GDC using the your dbGaP credentials.
https://gdc.cancer.gov/access-data/obtaining-access-controlled-data
Download Client Operation
Once you or your project leader have attained access to the reseach project,you will need to download access tokens from the gdc data portal. A comprehensive guide on how to use the GDC client is available here.
PDC
The Bionimbus Protected Data Cloud (PDC) is a secure biomedical cloud operated at FISMA moderate as IaaS with an NIH Trusted Partner status for analyzing and sharing protected datasets. The Bionimbus PDC is a collaboration between the University of Chicago Center for Data Intensive Science (CDIS) and the Open Commons Consortium (OCC). The Bionimbus PDC allows users authorized by NIH to compute over human genomic data in a secure compliant fashion.
It is a secure data cloud that stores US PCAWG data.
| Property | Value |
|---|---|
| Name | Bionimbus Protected Data Cloud |
| Contact | support@opensciencedatacloud.org |
| Repository type | Cloud |
| Official Website | https://bionimbus-pdc.opensciencedatacloud.org |
| ICGC Portal Page | Portal |
| Download Client | Amazon Web Services Command Line Interface |
| Client Documentation | AWS Guide |
| Repo Code | pdc |
Obtaining Data Access
Same as obtaining Data Access to GDC
Download Client Operation
The data in the PDC can be accessed using the AWS CLI. You will first need to enter your key and secret key with aws configure and follow the prompts. This key can be download from the projects tab of the official PDC website. Once your credentials have been entered, you can begin downloading.
Download manifest file from ICGC Portal
As described in the Search for PCAWG data section, once you satisfied with the search result of TCGA data files, click on the "Download Manifest" button as illustrated below to retrieve the manifest tarball (named as manifest.*.tar.gz). Unpack the tarbal, you should get a file named as manifest.pdc.*.sh, eg, manifest.pdc.1586448715169.sh.
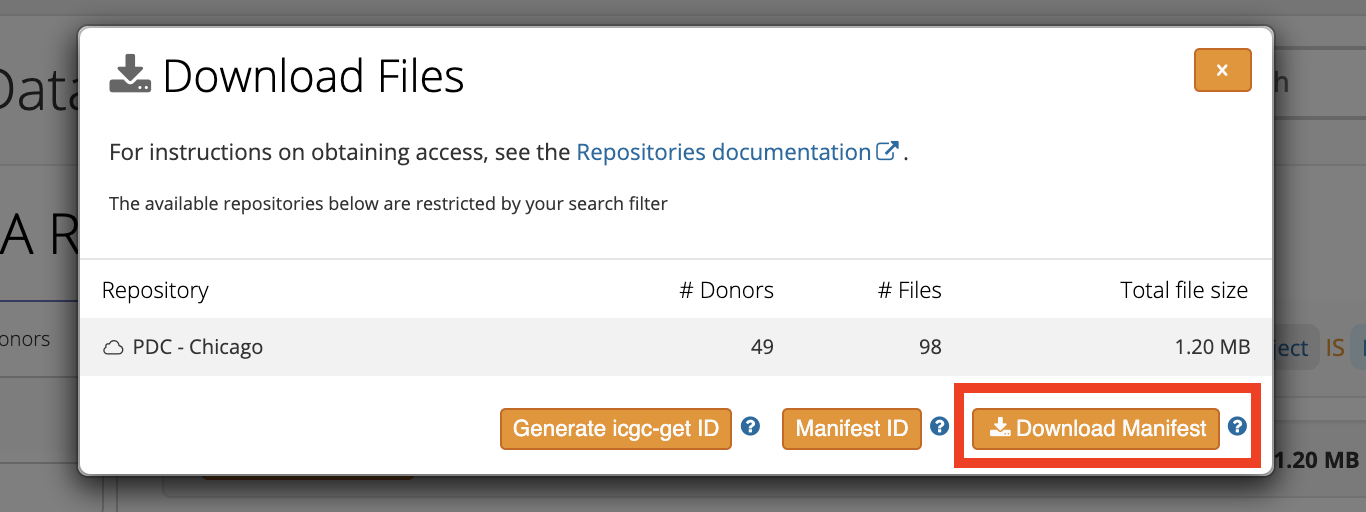
Convert ICGC manifest file to PDC's Gen3 manifest file
We need to convert the mainfest file to PDC's Gen3 manifest file before downloading the actual data files. A Python script (dcc_to_gen3.py) is needed to perform the conversion, the script can be downloaded with the following command:
wget https://raw.githubusercontent.com/uc-cdis/pdc_tools/1.0/dcc_manifest_conversion/dcc_to_gen3.py
You need to have Python 3 and required libraries (such as numpy and pandas) installed. Once installed, you can run the script to get Gen3 manifest file. Remember to replace the ICGC manifest with your own file name.
python dcc_to_gen3.py --manifest manifest.pdc.1586448715169.sh
This will produce a Gen3 manifest file named as gen3_manifest_manifest.pdc.1586448715169.sh.json, which contains information needed to download the acctual data from PDC using gen3-client tool.
Install Gen3-client
Run the following commands to install gen3-client if you are using macOS:
mkdir -p ~/.gen3
echo "" >> ~/.bashrc
echo "export PATH=\$PATH:~/.gen3" >> ~/.bashrc
curl https://api.github.com/repos/uc-cdis/cdis-data-client/releases/latest | grep browser_download_url.*osx | cut -d '"' -f 4 | wget -qi -
unzip dataclient_osx.zip
mv gen3-client ~/.gen3
rm dataclient_osx.zip
source ~/.bashrc
With that you should be able to run gen3-client command from your console and see the usage message.
For installing gen3-client on other OS, please follow instructions here.
Get gen3-client API key and configure your profile
Now you need to create gen3-client API key from https://icgc.bionimbus.org after authentication
via NIH eRA commons. To do that goto login page, and click on "Login with NIH" button. After authenticated successfully, please goto https://icgc.bionimbus.org/identity to create
the API key. On the popup dialog click on "Download json" to retrive API key, as shown below:
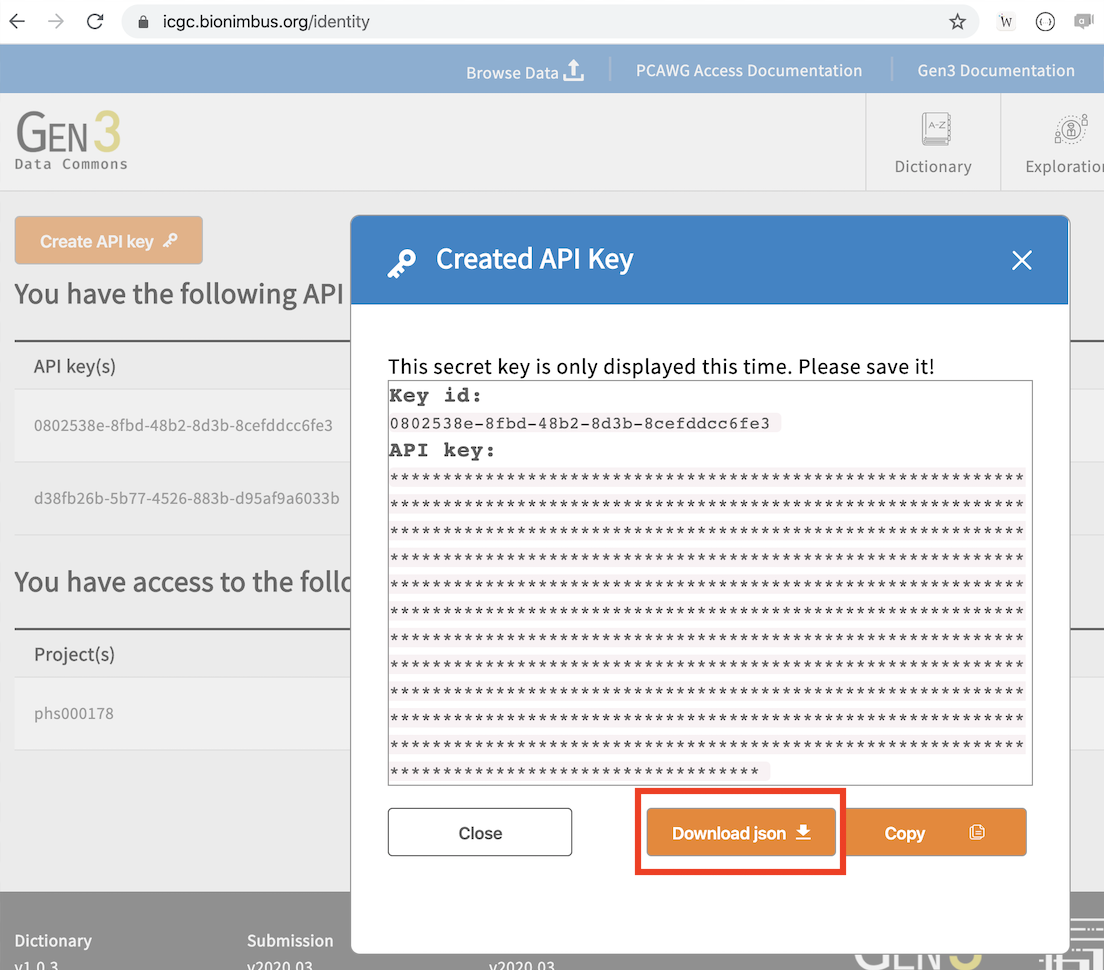
The API key will be saved as credentials.json. You can then use it to configure a profile, let's name the profile icgc:
gen3-client configure --profile=icgc --cred=credentials.json --apiendpoint=https://icgc.bionimbus.org/
Upon success, you should see a message: Profile 'icgc' has been configured successfully.
Download data using gen3-client download-multiple command
With icgc profile configured, you can download the PCAWG data using the gen3 manifest prepared earlier as follow:
gen3-client download-multiple --profile=icgc --manifest=gen3_manifest_manifest.pdc.1586448715169.sh.json --no-prompt
Azure
Microsoft Azure cloud service containing ICGC data.
| Property | Value |
|---|---|
| Name | Microsoft Azure |
| Contact | dcc-support@icgc.org |
| Repository type | Cloud |
| ICGC Portal Page | Portal |
| Download Client | Tarball, Docker |
| Repo Code | azure |
Obtaining Data Access
Follow the procedure outlined at the DACO page.
Download Client Operation
To operate the Score download client, follow the instructions here.